INTRODUCTION
Mangrove forests are highly productive ecosystems of intertidal areas in tropical and subtropical regions (Spalding et al., 2010; Tomlinson, 2016). Colombia is the third-largest mangrove country in the Latin America, with an approximate area of 371,081 ha. of which 282,835 ha. are on the Pacific coast (Giri et al., 2011).
In the Colombian Pacific, mangrove ecosystems consist mainly of eight species including Avicennia germinans (L.), Conocarpus erectus L., Laguncularia racemosa (L.) C.F. Gaertn., Mora oleifera (Triana ex Hemsl.) Ducke, Pelliciera rhizophorae Planch. and Triana, Rhizophora harrisonii Leechm., Rhizophora mangle L., and Rhizophora racemosa G. Mey. (García, 2010; Cornejo et al., 2014; Mejía et al., 2014). In addition, these species also provide important services to the local communities of the region (Palacios and Cantera, 2017). However, despite these benefits, combinations of human activities and natural events have been widely recognized to be associated with the decline of these trees in this part of the country (Prahl, 1989; Álvarez, 2003; López et al., 2016).
In recent years, the productive potential of mangrove ecosystems has been largely considered due to the metabolic activity of fungal communities (Hyde and Lee, 1995; Rineau et al., 2013; Thatoi et al., 2013). This potential has aroused great interest in investigating associations between mangrove trees and potentially beneficial fungi such as endophytes (Petrini, 1991; Wilson, 1995), which colonize the internal tissues of plants by protecting them from adverse factors (Gilbert et al., 2002; Arnold, 2007), or turning as opportunistic pathogens when plants are under different levels of stress (Slippers and Wingfield, 2007).
Numerous species of endophytic fungi have been reported in Colombia, either from native or non-native symptomatic or asymptomatic plants (e.g., Salazar and Garcia, 2005; Ordóez et al., 2012; Gamboa and Otero, 2016). However, studies on these fungal groups associated with mangrove species in the country remain very limited (Bolívar-Anillo et al., 2015). The objective of this study was to evaluate the presence of fungal endophytes associated with mangrove species in the San Pedro Nature Reserve, Buenaventura, Colombia, and to determine their possible role in disease development based on relevant literature.
STUDY AREA
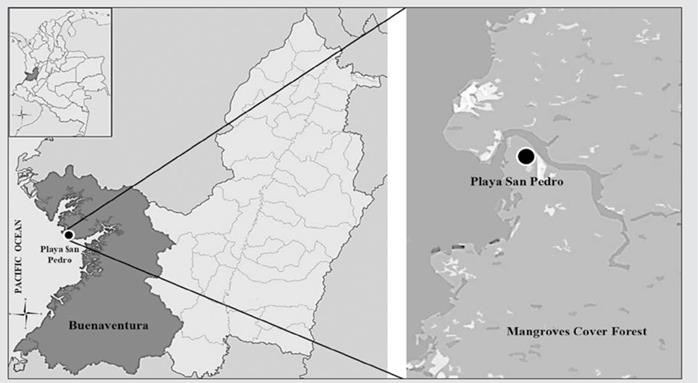
The San Pedro Nature Reserve (03° 50’ 06” N y 77° 15’ 28” W), is located at the isthmus of Pichidó, in the middle area of the Colombian Pacific coast, in the municipality of Buenaventura (Valle del Cauca). The reserve has an area of 100 ha, of which 40 % is mangrove ecosystem and 60 % rainforest. It has an average annual temperature of 26 °C, the relative humidity is 80 % to 98 %, and an average annual rainfall of 7,000 mm. Four mangrove species are present at this nature reserve, including Laguncularia racemosa, Mora oleifera, Pelliciera rhizophorae, and Rhizophora racemosa (Fig. 1).
MATERIALS AND METHODS
Field sampling
Prior recognition of the study area was performed, followed by a random sampling of asymptomatic branch segments from the middle part of canopy. In total 20 branches of approximately 10 cm long (ca.) were collected, five from each mangrove species including Laguncularia racemosa, Mora oleifera, Pelliciera rhizophorae and Rhizophora racemosa were placed in paper bags and stored in a portable refrigerator. They were subsequently transferred and preserved in the laboratory of the Biology Program of the Universidad del Quindío for further fungal isolation. It needs to be considered that the sampling size (number of branches collected) performed in this study depended on the sampling permits obtain from the National Environmental Licensing Authority.
Fungal isolations
The isolation of endophytes was carried out following the protocol described by Slippers and Wingfield (2007), with certain modifications made by Osorio et al., (2017). The collected branches were washed with tap water to remove excess debris and for each branch, five portions of 0.5 cm approximately were cut. These fragments were submerged in 90 % ethanol for one minute, followed by sodium hypochlorite (NaOCl) at 4 % (1 min), then in 70 % ethanol (1 min), and ending with a rinse in distilled water (1 min).
After surface sterilization and the pieces were placed on sterile paper for drying, each fragment was transferred to 2 % malt extract agar (MEA; 20 g malt extract, 20 g Biolab Agar in 1 L of distilled water), containing 50 mg of streptomycin to prevent bacterial colonization. The samples were incubated at room temperature, in natural light and darkness. The resulting colonies were purified using the “single hyphal tip” method, as described in the procedure carried out by Osorio et al., (2017), subsequently, the resulting colonies were separated into morph-groups for the following DNA extraction, which was performed by following the methodology of Raeder and Broda (1985).
Direct PCR amplification and sequencing
PCR was performed directly from representative pure cultures without prior DNA purification using Phire Plant Direct PCR kit (Thermo Scientific, USA) following the manufacturer’s instructions. ITS1-F and ITS4 primers (White et al., 1990) were used for the amplification of the ITS region. 25 µl PCR reactions contained 0.5 µl DNA template, 0.5 µM ITS1-F and ITS4 primers, 15 µl 2 × Phire Plant PCR buffer (includes dNTPs and MgCl2) (Thermo Scientific) and 0.5 µl Phire Hot Start II DNA Polymerase (Thermo Scientific). Initial denaturation was performed at 98 °C for 5 min followed by 35 cycles of (1) denaturation at 98 °C for 5 s, (2) annealing at 55 °C for 5 s, and (3) extension at 72 °C for 20 s. The final extension occurred at 72 °C for 1 min. The amplified PCR products were run on 1 % agarose gels, and successfully amplified products were purified with EXO-SAP protocol: the PCR product (20 μl) was mixed with 8 μl of Exo-SAP (5 μl of Exonuclease I (20 U/μl) (Fermentas,Vilnius, Lithuania) and 100 μl of Shrimp Alkaline Phosphatase (1 U/μL) (Roche Diagnostics, Indianapolis, USA) in a 1000 μl reaction mixture). The samples were incubated at (1) 37 °C for 15 minutes and (2) at 80 °C for 15 minutes. The PCR products were sequenced at Macrogen Europe. The resulting sequences were manually analyzed, and consensus sequences obtained by using Sequencher v.5 software.
Phylogenetic analysis
The obtained sequences were compared to the species data in GenBank (https://blast.ncbi.nlm.nih.gov/Blast.cgi) by performing BLAST searches on GenBank using the BLASTn algorithm (Altschul et al., 1990). DNA sequences of closely related species were obtained from GenBank and combined into datasets with the ITS sequences of the isolates obtained in this study (Table 1). The selection of the outgroup for each dataset was made according to the results of previous studies, including those of Sun and Guo (2012) and Osorio et al. (2017). The data sets were aligned using the MAFFT program (/), selecting the iterative refinement method G-INS-i (Kuraku et al., 2013; Katoh et al., 2019) and were manually edited with MEGA v. 10.1.7 (Kumar et al., 2018). Phylogenetic reconstructions were performed by maximum parsimony (MP), maximum likelihood (ML), and bayesian inference (BI) for each dataset obtained.
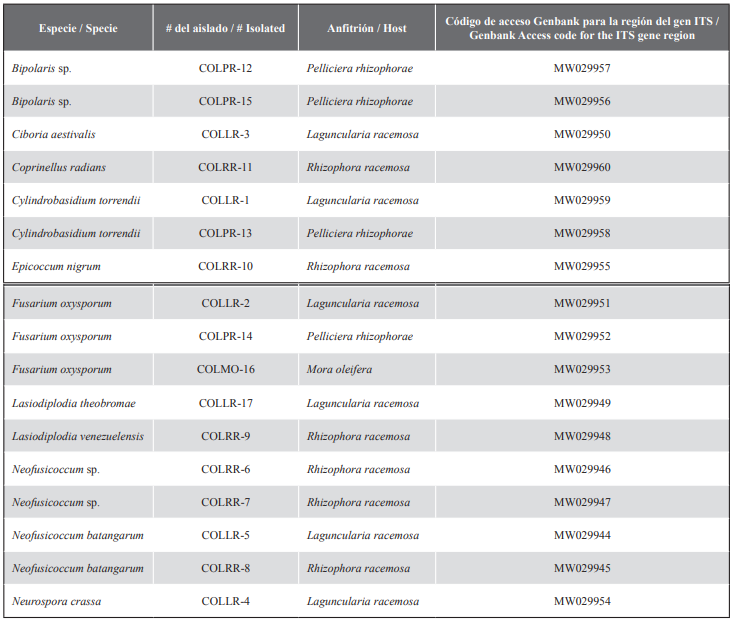
Table 1 Isolate numbers and access information to GenBank of isolates obtained from mangrove trees in the San Pedro Buenaventura Nature Reserve (Colombia).
Analyses of the MP were performed in PAUP v. 4. 0b10 (Swofford, 2003) using a heuristic search and tree-bisection-reconnection (TBR) reorganization algorithm. Gaps were treated as missing data and all characters were treated as unordered type with the same weight. Statistical support for the nodes was obtained by performing 1000 bootstrap replicates. For ML and BI analyses, replacement models were previously obtained using the jModelTest software v. 2.1.10 (Darriba et al., 2012) using the Akaike (AIC) information criteria (Akaike, 1974). ML analyses were performed with the PhyML v. 3.1 program (Guindon and Gascuel, 2003), considering the models obtained with jModelTest. The support values for the nodes were estimated using 1000 ML bootstrap replicates. Subsequent probabilities for BI were determined based on the Markov Chain Monte Carlo (MCMC) algorithm that was performed on MrBayes v. 3.2.7a (Ronquist et al., 2012). Two independent runs were conducted simultaneously for 5 million generations. The burn-in values were determined through Tracer software v. 1.7.1 (Rambaut et al., 2018), the first 0.25 sampled trees that represented the burn-in phase were discarded and the remaining trees were used to build the consensus tree. Finally, the phylogenetic trees obtained from the BI, ML and MP analyses were edited in FigTree v. 1.4.4 (http://tree.bio.ed.ac.uk/software/figtree/) and/or MEGA v. 10.1.7 (Kumar et al., 2018).
RESULTS
A total of 20 branches were sampled, belonging to four mangrove species. Of these, 17 endophytic fungi were isolated, representing nine genera belonging to eight families including Botryosphaeriaceae, Didymellaceae, Nectriaceae, Physalacriaceae, Pleosporaceae, Psathyrellaceae, Sclerotiniaceae and Sordariaceae; two in the Basidiomycota and six in the Ascomycota. Of these isolates, six were obtained from L. racemosa, six from R. racemosa, four from P. rhizophorae and one from M. oleifera.
PCR amplification and sequencing
Fragments obtained from the ITS sequences, comprised approximately 465 to 606 bp in size. The sequences included in this study were deposited in the GenBank DNA sequence database (Table 1).
Phylogenetic analysis
Analyses of BI, ML and MP, for the ITS gene region showed congruent topologies supported by bootstrap support values and subsequent probabilities, thus identifying eight fungal families (Fig. 2). The phylogeny generated for Botryosphaeriaceae included fifteen species previously described based on the ITS region (Fig. 3a). In Didymellaceae 18 species were phylogenetically compared, of which only one isolate (MW029955), was obtained from mangroves (Fig. 3b). For Nectriaceae, fourteen species described earlier were used as references; three isolates obtained from mangrove trees (MW029951, MW029952, MW029953) grouped in the same clade with Fusarium falciforme (Carrión) Summerb. and Schroers 2002, F. oxysporum and F. solani (Fig. 4a).
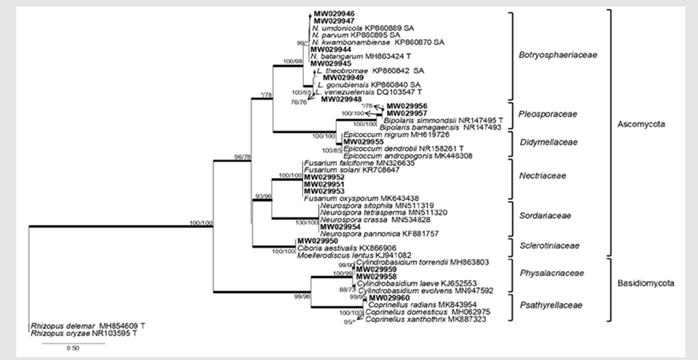
Figure 2 Families of isolated endophyte fungi of four mangrove species. A phylogram obtained from the BI, ML and MP analyses of the ITS dataset. The sequences obtained in this study in bold. The subsequent probabilities ≥ 95 % of BI are represented by thick branches. Bootstrap support values > 70 % are indicated near nodes as MP/ML. * • Bootstrap support values < 70 %.
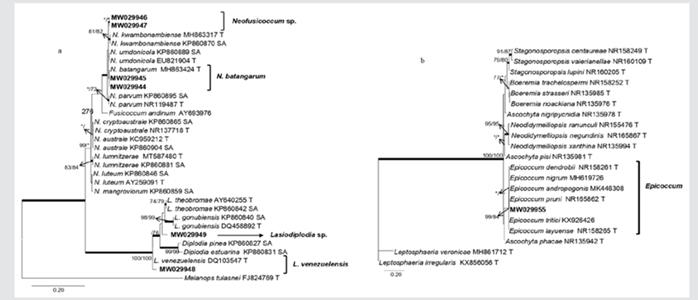
Figure 3 Phylogram of genera in Botryosphaeriaceae. Six of the 17 mangrove isolates are closely grouped with the genera Lasiodiplodia and Neofusicoccum (3a). A phylogram of Didymellaceae genera, indicating that one of the isolates obtained from Rhizophora racemosa (in bold) is grouped with different species of Epicoccum (3b). Phylograms obtained from BI, ML and MP analyses of the ITS dataset. The subsequent probabilities ≥ 95 % of IB are represented by thick branches. Bootstrap support values > 70 % are indicated near nodes as MP/ML. * • Bootstrap support values < 70 %.
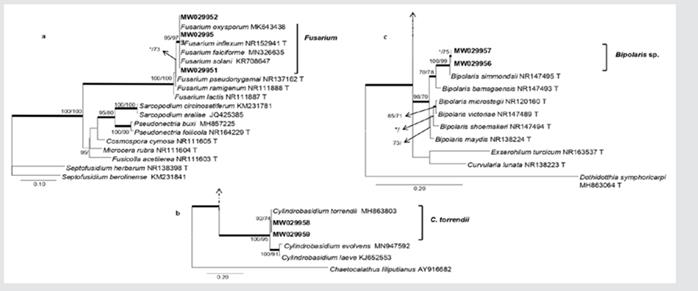
Figure 4 Phylogenetic analysis of genera in Nectriaceae. Mangrove isolates (in bold) are grouped with different species of Fusarium (4a). Phylogram with genus Physalacriaceae, the isolates obtained from mangroves are clearly grouping with Cylindrobasidium torrendii (4b). Phylogram of Pleosporaceae genera. Isolations obtained from Pelliciera rhizophorae (in bold) suggest a close relationship with Bipolaris simmondsii (4c). Phylograms obtained from BI, ML and MP analyses of the ITS dataset. The subsequent probabilities ≥ 95 % of IB are represented by thick branches. Bootstrap support values > 70 % are indicated near nodes as MP/ML. * • Bootstrap support values < 70 %.
Phylogeny of Physalacriaceae included nine previously described species; two isolates obtained from mangroves (MW029959, MW029958) were closely related to Cylindrobasidium torrendii (Fig. 4b). In Pleosporaceae, 16 species described earlier were included in the phylogenetic analysis; two isolates obtained from mangroves (MW029957, MW029956) formed a separate clade within the family (Fig. 4c). In Psathyrellaceae, 13 species were included in the phylogenetic analyses, of these, a single isolate obtained from mangroves (MW029960) grouped with Coprinellus radians (Fig. 5a). In Sclerotiniaceae, the isolate (MW029950) was closely grouping with two species in the same clade, Ciboria aestivalis and Moellerodiscus lentus (Fig. 5b). In Sordariaceae, a mangrove isolate (MW029954) was grouping with Neurospora crassa, N. pannonica, N. sitophila and N. tetrasperma (Fig. 5c).
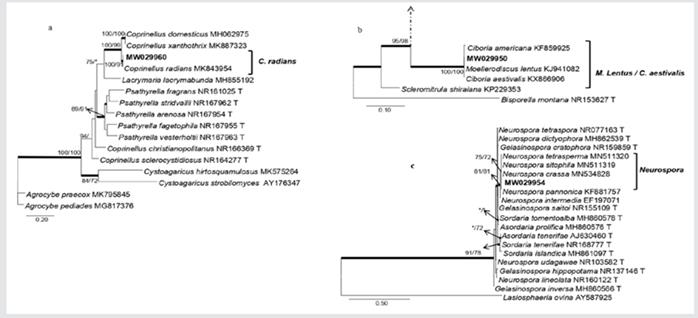
Figure 5 Phylograms obtained from genera in Psathyrellaceae indicating that one of the isolations obtained from Rhizophora racemosa (in bold) is closely related to Coprinellus radians (5a). Phylogram of genera in Sclerotiniaceae indicating that one of the isolates obtained from Laguncularia racemosa (in bold) groups with Moellerodiscus lentus and Ciboria aestivalis (5b). Phylogram obtained from genera in Sordariaceae,shows that isolations obtained from branches of Laguncularia racemosa (in bold) groups with four species of Neurospora (5c). Phylograms obtained from BI, ML and MP analyses of the ITS dataset. The subsequent probabilities ≥ 95 % of IB are represented by thick branches. Bootstrap support values > 70 % are indicated near nodes as MP/ML. * • Bootstrap support values < 70 %.
DISCUSSION
This study represents the first phylogenetic approach of endophytic fungi associated with mangrove trees in an area of the Colombian Pacific coast. Nine fungal genera were identified based on the DNA sequence data for the ITS region, a gene region widely used as a molecular marker in fungal identification and taxonomic studies (Hibbett et al., 2011; Joel and Bhimba, 2013). Based on the phylogenetic analysis, the identity of the isolates were verified and revealed that the endophytes collected in this study closely grouped with species such as Ciboria aestivalis, Coprinellus radians, Cylindrobasidium torrendii, Epicoccum nigrum, Fusarium oxysporum, Lasiodiplodia theobromae, Lasiodiplodia venezuelensis, Neofusicoccum batangarum and Neurospora crassa, and two isolates that could not be confirmed to species level, Bipolaris sp. and Neofusicoccum sp., all are reported for the first time at the Colombian Pacific coast.
The fungi obtained in this study belong mostly to Ascomycota, with fourteen isolates within seven genera; meanwhile, three isolates were members of Basidiomycota. Interestingly, previous studies have confirmed that the aerial and submerge parts of mangrove trees, are largely dominated by ascomycetes, while basidiomycetes have a lower proportion (Alias et al., 2010; Sun and Guo, 2012; Abdel et al., 2014; Simões et al., 2015), this is consistent with the saprophytic activity that present a large number of Basidiomycetes (Agrios, 2005; Mohapatra, 2008). While most Basidiomycetes are mostly found in terrestrial environments (Shearer et al., 2007), the Ascomycetes are considered as an important ecological group, with a wide variety occurring in diverse environments, including marine environments (Jones and Tan, 1987; Hyde and Jones, 1988; Abdel-Wahab, 2005; Vittal and Sarma, 2006; Alias et al., 2010; Suetrong et al., 2017). Moreover, a wide range of variables including host age, architecture, brightness levels, genotype, height of crown, among others, have been considered to affect fungal community structure, as well as endophyte-plant interactions (Saikkonen, 2007).
Two genera in the Botryosphaeriaceae, including Lasiodiplodia and Neofusicoccum were obtained in this study, it is consistent with previous studies reporting their wide range of ecologically diverse fungi, found in all geographical and climatic areas of the world (except in polar regions) (Slippers and Wingfield, 2007; Phillips et al., 2013). For instance, reports of Botryosphaeria, Lasiodiplodia, and Neofusicoccum in mangroves of Brazil (De Souza et al., 2013), Neofusicoccum in China (Xing et al., 2011) and fourteen species belonging to four genera, Botryosphaeria, Diplodia, Lasiodiplodia, and Neofusicoccum in mangroves of South Africa (Osorio et al., 2017), confirms the ubiquity of the species belonging to this fungal family.
Neofusicoccum was the most commonly isolated genus, given the ability of species of this genus to inhabit a wide variety of plant species and geographical areas worldwide (Slippers et al., 2005; Sakalidis et al., 2011). This is consistent with our results, since Neofusicoccum was the most isolated genus, indicating its wide geographical range (Shetty et al., 2011).
Lasiodiplodia theobromae is considered a highly phytopathogenic fungus (Slippers and Wingfield, 2007), which is commonly found in tropical and subtropical regions (Punithalingam, 1980; Burgess et al., 2006; Phillips et al., 2013; Mehl et al., 2017). Previous reports indicate that this species is associated with different host plants, causing plan diseases and being responsible for serious damage to crops of economic importance (Coutinho et al., 2017). It has also been reported as the causal agent of rot, withering and necrosis of seeds in mangrove plants in India (Misral, 2002), and also as a fungal pathogen of mangrove plants in the Niger Delta in Nigeria (Ukoima and Amakiri 2000), possibly in response to an imbalance in the mangrove ecosystems. It is then of concern, considering that mangroves in in Buenaventura are under constant stress due to the anthropogenic disturbances observed during field work, although no obvious symptoms of disease were found. In this sense, further studies are highly recommended to determine the degree of pathogenicity of this species in Colombian mangroves.
In this study, phylogenetic analyses supported that the isolate MW029948 is closely related to Lasiodiplodia venezuelensis, isolated from branches of R. racemosa. To date, this species has not been widely reported in other parts of the world; L. venezuelensis is considered as a rare species having geographically limited distribution (Mohali, et al., 2017). This is the first report of L. venezuelensis associated with mangrove trees in Colombia.
The isolates MW029951, MW029952 and MW029953 grouped within the genus Fusarium, and subsequent analyses revealed their proximity to F. oxysporum, which is a widely existing species (Munkvold, 2017). This isolate was obtained from three of the four mangroves sampled in San Pedro; L. racemosa, M. oleifera and P. rhizophorae, and was the only isolated species from M. oleifera. The genus includes numerous ubiquitous species that can colonize a wide range of habitats (Munkvold, 2017). In mangrove environments the genus has been found colonizing the mangroves of Mangalvan (Pal and Purkayastha, 1992), the leaves and branches of Avicennia schaueriana, L. racemosa and R. mangle in Brazil (De Souza et al., 2013), and recently in microbiome of the mangroves rhizosphere of Avicennia marina in the Red Sea, Saudi Arabia (Simões et al., 2015). Also, members of the F. oxysporum complex are known to cause vascular withering diseases, and are important as root rot pathogens (Munkvold, 2017). Although F. oxysporum includes both pathogenic and nonpathogenic strains, it will be then important to conduct future studies to investigate its pathogenicity in mangroves, and thus to estimate a possible threat to these ecosystems.
Members of the Didymellaceae are cosmopolitan (Aveskamp et al., 2008, 2009; Chen et al., 2015a, 2015b). The isolate MW029955 resides in this family, being phylogenetically closely related to Epicoccum nigrum (99 / 100 %). Different species of Epicoccum are known to produce bioactive secondary metabolites with antiviral (Guo et al., 2009), antifungals (Harwoko et al., 2019), anti-inflammatory (Gonda et al., 2016), and cytotoxic activities (Wang et al., 2010). Among these species, the E. nigrum species complex (Favaro et al., 2011), has been an important part of research on bioactive compounds of endophytic fungi isolated from the mangrove Acanthus ilicifolius L. (Yan et al., 2019). Such research is also facilitated due to the wide geographical distribution and high hosts’ variability (Perveen et al., 2017; Ahumada et al., 2018; Rhim et al., 2019; El-Sayed etal., 2020).
The current phylogenetic analyses confirmed that the isolate obtained from R. racemosa (MW029960) corresponded to Coprinellus radians. Coprinellus is one of the most important lineages of Coprinoid fungi in the Psathyrellaceae (Hussain et al., 2018), with approximately 80 described species (Kirk et al., 2008; Nagy et al., 2012; Gomes and Wartchow, 2014). These fungi are common saprotrophs in wood chips, leaf blades and herbivorous manure (Schafer, 2010). Coprinellus radians has been mainly isolated from human patients (Lu et al., 2020), however, information on its relationship to mangrove trees still remains very limited. Neurospora crassa isolated from branches of L. racemosa in San Pedro, was also isolated from leaves of Rhizophora mucronata Lam (Joel and Bhimba, 2013) to determine their antimicrobial and cytotoxic activity. Neurospora is common in humid tropical and subtropical regions (Turner et al., 2001), also found in mangrove sediments in India (Thamizhmani and Senthilkumaran, 2012) and in the mycobiome of the rhizosphere of the red Sea marine Avicennia mangroves (Simões et al., 2015). Both C. radians and N. crassa are species that represent a promising source of bioactive molecules with potential use in the pharmaceutical industry (Demers et al., 2018).
Of the four mangrove species sampled in San Pedro, L. racemosa and R. racemosa have previously been reported as hosts of endophytic fungi (Vázquez et al., 2000; De Souza et al., 2013; Ukoima et al., 2013), while for M. oleifera and P. rhizophorae there is still no data available. Likewise, information for Bipolaris sp., Ciboria aestivalis and Cylindrobasidium torrendii known to be associated with mangrove trees remain limited. Regarding M. oleifera and P. rhizophorae, the former belongs to a genus native to lowlands of the rainforests of South America, Central America and the Caribbean islands (Bernal et al., 2015), and the latter is the only species of the genus, being native to Central America from Honduras to Ecuador (Ulloa et al., 2017). This limited distribution of hosts, coupled with the limited fungal studies associated with mangrove ecosystems, and illustrates the lack of information on fungal microorganisms associated with these trees worldwide. Bipolaris (MW029956 and MW029957) includes members considered important pathogens of more than 60 plant genera (Sivanesan, 1984; Agrios, 2005; Manamgoda et al., 2011; Hyde et al., 2014).
Phylogenetic analyses also confirmed that the isolate MW029950 obtained from L. racemosa was closely relate to C. aestivalis. This species has been found in herbarium specimens of Cydonia oblonga Miller (Rosaceae) (Galan and Palmer, 2001). In addition, current analyses supported that the isolates MW029958 and MW029959 were closely related to C. torrendii, a fungus that causes weak and limited decomposition of wood (Floudas et al., 2015). These fungal groups are reported for the first time from asymptomatic branches of mangroves in Colombia.
Despite Colombia’s great floristic richness, studies in terms of its associated mycobiota remain very limited. In this sense, this study contributes to the knowledge of fungal diversity associated with mangrove trees in Colombia.











 text in
text in