INTRODUCTION
Allelopathy is a process in which plants release into the environment chemical compounds, which suppress, inhibit, or displace the establishment or growth of other plants in their habitat [1, 2]. The allelopathic potential of some species of crops and weeds influence growth and distribution of some of the weed species associated to yield of desired plants, which is why allelopathy is successfully employed in biological control programs centered around phytopathology and problematic weed control. In every allelopathic phenomenon there is a donor plant that releases chemical substances into the environment by a certain route (e.g. lixiviation, residue decomposition, etc.), which cause a synergistic or antagonistic effect on germination physiology, growth, or plant development when incorporated by a receptor plant. Allelopathic compounds that unleash this process are called allelopathic compounds, agents, or substances. There are two types of allelopathy: eccrisodynamia, the influence by metabolites from autotrophic organisms, and saproccrinodynamia, the influence by metabolites produced by saprophyte organisms. Most allelopathic compounds released by plants are considered secondary metabolites produced as a result of primary metabolic pathways [3]. Depending on their phytotoxic action, bioactive concentration, persistence, and fate in the environment in which they are released, they can act as allelopathic compounds [4]. In addition, the allelopathic effect in nature is originated by the joint action of several allelochemicals, instead of by the action of only one [5]. It is possible to extract three important and essential features when studying the existence of allelopathic phenomena: 1) the release into the environment of a compound in charge of transmitting an effect, 2) its absorption by the receptor organism, and 3) an effect on growth. The complexity of the phenomenon results from the variables in each of these processes, for example, the mechanism of allelochemical release and its stability in the environment, which will define the levels and the manner in which it will be absorbed by the receptor mechanism, and the uncountable ways of action found in nature, which in many cases depend on the receptor being.
On the other hand, metabolomics studies the complete set of metabolites in an organism [6]. Metabolomics analyzes a wide range of different compounds, which differ in many aspects: chemical nature, solubility and concentration [7]. Metabolomics allows for the analysis of several chemical compounds contained in plant cells that contain a great number of metabolites of low molecular weight (<1.500 Da) [8]. Today, metabolomics have to a large extent expanded metabolite profile, fingerprint collection, and selection and identification of metabolite markers. Thus, the present review is focused on providing a recent update on the science of allelopathy in the context of physiology and practical application of metabolomics in the analysis of the allelopathome of plants with economic interest and finally mentionated the biological activity of allelopathic secondary metabolites.
ALLELOPATHY
The term allelopathy (from the Greek allelon: one to another andpathos:to suffer; a harmful effect of one to another) was initially used for the first time to refer to the harmful of beneficial effects that are direct or indirect result of the action of chemical compounds released by a plant, exerting its action on another one. Allelopathy is a form of interference caused by plant growth as a result of chemical interactions between plants (through living or decomposing tissue) and other organisms mediated by the release of bioactive secondary metabolites produced by the plant, called allelochemicals (chemical inhibitors) [9]. Such biochemical interactions can have inhibitory or stimulating actions. Allelopathy is present in many plant communities, in the ecological process, these interactions are of great importance for natural ecosystems and for those that show some type of management, due to their participation in the process of succession and structure, composition, and dynamics of native or cultivated plant communities [10]. Due to the way of action of these compounds, they become an alternative as herbicides, insecticides, or nematicides. Allelopathy is used to suppress weeds in crop fields. Examples of important allelopathic crops are alfalfa, asparagus, rye, sorghum, rice, sunflower, sugar cane, rapeseed, wheat, among others [11].
Allelochemicals are found in pilose roots, sprout tissues, aqueous extracts, dried residues, decomposing leaves, flowers, branches, root exudates, and straw [12]. Allelopathic effects are modifiable by additional biotic and abiotic stress factors, uncertain meteorological incidents, or physical, chemical, and biological soil factors, which can influence residence time, persistence, concentration, and the fate of allelopathic compounds in the environment. In allelopathy, there is a plant that produces the allelo-pathic agent and a receptor plant from another species. When the producer and the receiver plant are the same species, it can be considered as a special allelopathy case called autotoxicity. Allelopathic activity depends on several factors, such as 1) sensitivity of receiver species, 2) release of the toxin into the environment, and 3) activity and biotic and abiotic interactions occurring in the soil with the toxin (microorganisms, pH, temperature) [13].
It is necessary to establish four conditions for an interaction to be considered as allelopathy: 1) to prove the existence of interferences, to describe the symptoms and to quantify the degree of interference, 2) to isolate, assay, and characterize the allelochemicals, 3) interference symptoms previously diagnosed, must be repeated by the application of toxins in doses present in nature, and 4) the release of toxins, their movement and uptake by the receiver plant must be monitored and it must be proved that the dose is high enough to explain the observed interference. In the testing assays for the phenomenon of allelopathy, called bioassays, it is necessary and of great importance to control the parameters such as temperature and water availability, in order to decrease the number of factors that can influence it, which allows for research on the action of other mechanisms [14]. In germination, allelopathy cannot exactly act on germinality (final germination potential with time), but on germination velocity. Alterations on the germination pattern can occur because of membrane permeability, DNA transcription and translation, secondary messenger action, respiration, enzyme and receptor conformation, and/or the combination of these factors.
Allelopathic mechanisms
The study of the plant secondary metabolism reveals that plants produce a wide range of substances that, in addition to performing physiological functions, also allow for the interaction among individuals, which impacts on the environment. These chemical substances, called allelochemicals, are studied by allelochemy or allelochemistry, which contribute to species adaptation and participate in the organization of plant communities [14]. Most allelochemicals are autotoxic, but plants have developed methods to store them without being affected. Many are located inside certain tissues, cells, or organelles, with the purpose of isolating them from critical metabolic processes in the organism that produces them. In other cases, toxins are accumulated in an active way and are chemically transformed before their release. Several mechanisms for the release of allelochemicals have been studied in several plant tissues, including volatilization or lixiviation of aerial parts, root exudation, and vegetal residues decomposition in soil (Latif et al., 2017) [9]. Radicular exudation are those organic compounds released into the environment by the roots of healthy and intact plants. Phenolic compounds have been also been reported to be the compounds most commonly involved in allelochemicals and responsible for allelopathic activity [15].
Metabolomics
Metabolomics studies, identifies, and quantifies in a systematic manner the totality of metabolites in a biological system (cells, tissues, or biological fluids) that are product of metabolic reactions in the living beings [16], and reflect their physiological and biochemical states, derived from the transcriptome and proteome, with the joint action of post-translational environmental impact, enzymatic activity, and even membrane transport that are not always directly modulated by genic expression.
The fields of application of metabolomics are agriculture, pharmaceutical industry, environmental studies, and food industry. Metabolomics emerged as a tool in biological sciences to identify subtle changes in metabolic profiles between biological systems in different physiological or pathological states [16]. The metabolome can be understood as the final product of gene expression with a possible impact on cell, tissue, or organ-ismic phenotype, hence that quantification of metabolites provides a wide perspective of the biochemical status that can be used to monitor and establish gen function [17]. Depending on the field of study, metabolomics makes use of different strategies: 1) Target analysis: the objective is the quantitative analysis of the metabolites of the substrate and/or the product of a target protein, 2) Metabolic profiling: it is centered around the quantitative analysis of a set of compounds of a determined biosynthetic pathway or a group of related metabolites; footprinting: the analysis of (exo)metabolites secreted/excreted by an organism; if the organism is culture-grown, the analysis of environmental conditions and growth substances can be included; and 4) fingerprinting: the analysis of intracellular metabolites with biochemical substances to classify the sample according to its origin or status (case/control, healthy/sick) [17]. Metabolome research has gained sophistication by its huge complexity and dynamism. An ideal analytic technique should be performed directly on the samples, without the need of pretreatment. In addition, it should be reproducible, robust, sensitive, and precise [18]; however, there is no technique that can provide all these characteristics. The major analytic techniques employed in metabolomics are based on nuclear magnetic resonance (NMR) and mass spectrometry (MS). The latter has been established in recent years as an essential tool in metabolite analysis; it requires the pre-preparation of metabolic components, whether by gas chromatography (GC) or liquid chromatography (LC, HPLC). Capillary electrophoresis (CE) coupled to mass spectrometry (MS) is seen as a promising alternative, just like the Fourier Transform Infrared Spectrometry (FTIR). Enzymatic assays have also been used in metabolite determination [19].
Metabolomic studies in allelopathy
It has been showed that certain plants release products, called allelochemicals, in their immediate environment, affecting the growth and development of other neighboring species [20]. Allelopathic interactions play an important role in natural plant ecosystems. Metabolomics shows diverse applications such as the characterization of biochemical variations, the discovery of the potential for targets in metabolic engineering, the examination of responses of the plant to the environment and its responses to biotic and abiotic stressors. Metabolite concentration varies in response to genetic perturbation, elicitation, and environmental stimuli. According to the different structures and allelochemical links, these are classified in the following categories: 1) water soluble organic acids, linear chain alcohols, aliphatic aldehydes, and ketones, 2) simple unsaturated lactones, 3) long chain fatty acids and polyacetylenes, 4) quinines (benzo-quinone, antraquinone and quinine complexes), 5) phenolic compounds, 6) cinnamic acid and its derivatives, 7) coumarins, 8) flavonoids, 9) tannins, and 10) steroids and terpenoids (sesquiterpene, diterpene, and triterpene lactones) [21]. In the following lines some relevant studies regarding metabolomic studies in allelopathy of plants of agro-economic interest.
Allelochemicals analyzed by metabolomics in alfalfa
The importance of alfalfa (Medicago sativa L.) is because of its biological and pharmacological activity. It has antimicrobial, antifungal, antioxidant, anti-inflammatory, anticancer, and phytoestrogenic activity, it is a central nervous system anxiolytic, and it reduces cholesterol [22]. M. trunculata produces flavonoids and isoflavonoids in its roots and aerial parts [23].
M. sativa produces flavonoids, phenolic acids and saponins of horticultural and agricultural importance, which are used against a wide range of pests such as nematodes, herbivore insects and fungi [22]. The physiological effect of alfalfa reduces weed growth and agricultural crops through water-soluble allelochemicals [24].
Phenolic compounds produced by M. sativa are cinnamic acid, caffeic acid, vanillic acid, p-coumaric acid, hydroxybenzoic acid, transcinnamic acid, ferulic acid, and medi-carpin (figure 1).
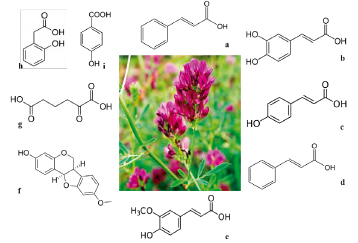
Source: own preparation.
Figure 1 Principal allelopathics metabolites ofM. sativa. a) Cinnamic acid (148.16 g/mol, C9H8O2), b) Caffeic acid (180.16 g/mol, C9H8O4), c) p-coumaric acid (164.05 g/mol, C9H8O3), d) Transcinnamic acid (148.16 g/mol, C9H8O2), e) Ferulic acid (194.18 g/mol, C10H10O4), f ) Medicarpin (270.27 g/mol, C16H14O4), g) 2-oxoadipate (160.125 g/mol, C6H8O5), h) 2-hydroxyphenylacetic acid (152.15 g/mol, HOC6H4CH2CO2H), and i) hydroxybenzoic acid (138.03 g/mol, C7H6O3).
Stochmal y Oleszek (2007) found different levels of flavones in ten varieties of alfalfa. Approaches to metabolomic studies in alfalfa are diverse [25]. Several authors have focused on studying metabolomics of M. sativa. Today, alfalfa metabolites have been analyzed in conditions of alkaline stress using Gas Chromatography combined with Time-of Flight Mass Spectrometry (GC/TOF-MS) [26]; the metabolomic relationship between decrease of nutritive value and biological processes has also been studied with the purpose of producing alfalfa hay [27]. In addition, changes presented by C and N in the primary metabolism of amino acids under water-limiting conditions in alfalfa have been studied [28]. Zubair et al. (2017) metabolomically characterized 5880 chemical compounds from root exudates between allelopathic genotypes of M. sativa [29]. A metabolomic analysis was carried out in root exudates of M. sativa using mass detection by Quadruple Time-of-Flight (Q-TOF). The work shows that inhibition of root and sprout growth of Lolium multiflorum was due to the allelopathic compounds released by the genetic variation of alfalfa genotypes. In the root exudates of the weakly allelopathic genotypes three metabolites were found: 2-oxoadipate, 2-hydroxyphenylacetic acid, and 2'-deoxyuridine 5'-mono-phosphate [29].
Allelochemicals analyzed by metabolomics in rice
Rice (Oryza sativa) is a food crop consumed in all the world. O. sativa has potential benefical effects on health related to the polyphenolic content of the grain, which prevents vents cancer, cardiovascular diseases, and diabetes complications [30]. Several types of secondary metabolites produced by rice have been identified using spectroscopy techniques. Pigmented rice contains a higher amount of phenolic compounds, sucrose hydroxycinnamate esters and flavonoids (water soluble metabolites), of the flavonol type, such as quercetin, kaempferol and apigenin [8] than non-pigmented rice. Rice produces several types of secondary metabolites to protect itself against biotic and abiotic stress [31]. Sakuranetin levels (phytoalexin), subtly detectable in healthy rice leaves, increased rapidly under biotic and abiotic stress stimuli, including UV treatment and pathogen attack [32].
Field and laboratory experiments in many countries mentioned that rice is allelopathic and release allelochemicals in its environment. Some important allelochemicals in rice are phenolic acids (p-coumaric, ferulic, chlorogenic, caffeic and sinapic acids), phenyl alkanoic acids, fatty acids (phosphatidylinositol), hydroxyamic acids, indoles, and terpenes, diterpenoid momilactones related to labdane are the most important ones, and momilactone B has a particularly essential role (figure 2).

Source: own preparation.
Figure 2 Principal allelopathics metabolites of O. sativa. a) Momilactone B (330.42 g/mol, C20H26O4), b) p-coumaric (164.05 g/mol, C9H8O3), c) ferulic acid (194.18 g/mol, C10H10O4), d) chlorogenic acid (354.09 g/mol, C16H18O9), e) caffeic acid (180.16 g/mol; C9H8O4), f) sinapic acid (224.21 g/mol, C11H12O5), and g) phosphatidylinositol (886.56 g/mol, C47H83O13P).
Rice plants secrete momilactone B from its roots into neighboring environments throughout all its life cycle at phytotoxic levels. In addition, genetic studies of the rice genome show that momilactone elimination from root radicular exudates decrease allelopathy, which shows that they serve as allelochemicals. The involvement of allelopathy plays an important ecological role in rice evolution [33]. Bran rice oil contains nutritional and commercial phytochemical compounds, like carotenoids and g-oryz-anol (a mixture of ferulic acid with sterols and triterpene alcohols) [34]. Bran rice is the residue left after refining rice; it has a dark brown, red or purple color, and contains anthocyanins (flavonoids of reddish to purple color) and it is believed that they are bioactive components, for example, diet antioxidants, in said colored rice [8]. Metabolomic research in rice has been focused on the study of biotic stress (by pathogens), and abiotic stress of which different metabolite biomarkers associated with tolerance to temperature-generated stress, saline stress, drought, ozone [31] genetic modification (enzymatic expression, Na+ transporters), natural variation (genes), elucidation of systems for regulating metabolomic networks (study of primary metabolites) and nutritional starvation have been identified [35]. Matsuda et al. (2012) identified a key gene regulating the metabolism specialized in the flavonoids biosynthetic pathway [36]. The effective use of metabolomics and QTL analysis through the mapping of populations to delimit genetic control on rice metabolism has been reported [15, 37]. Metabolic approaches will lead the identification of secondary metabolites, and some of them could be used to improve rice quality and yield.
Allelochemicals analyzed by metabolomics in canola
Canola (Brassica napus) is the second oily seed most produced in the world after soy [38]. B. napus oil is largely composed of unsaturated long chain fatty acids such as oleic, linoleic and alpha-linolenic. These fatty acids show antibacterial activity [39, 40]. In addition, B. napus biosynthezes glucosinolates, phenylpropanoids, flavonoids (kaemp-ferol, quercetin, and isorhamnetin), and phytohormons (figure 3), these compounds have been identified by mass spectroscopy [41]. There is currently little information regarding the analysis of the metabolome of plant seeds. In the case of B. napus, from the metabolomic perspective, it is interesting to study seed metabolome to understand seed biology and physiology functions to better understand qualitative traits and products that can be obtained from seeds. Some studies have been carried out on A. thaliana [42], soy bean [43], pea [44] and tomato [45]. In a metabolomics study based on NMR, it was shown that unsaturated fatty acids, sucrose, and sinapine were the most discriminating metabolites between B. napus and B. rapa metabolites [46].

Source: own preparation.
Figure 3 Principal allelopathics metabolites of B. napus. a) synapil alcohol (210.23 g/mol, C11H14O4), b) p-hydroxybenzoic acid (138.03 g/mol, C7H6O3), c) quercetin (302.24 g/mol, C15H10O7), d) 3,5,6,7,8-pentahydroxyflavone(302.236g/mol, C15H10O7),e)sinapine(310.370g/mol, C16H24NO5), f) kaempferol (286.23 g/mol, C15H10O6), and g) isorhamnetin (316,26 g/mol, C16H12O7).
Asaduzzaman et al. (2015) carried out a metabolomic analysis and an allelopathic bioassay in order to compare three genotypes of allelopathic canola and three genotypes of non-allelopathic canola [47]. The results of the bioassay of the laboratory showed that there were differences between the genotypes in their capacity to inhibit the growth of ryegrass roots and sprouts. Root exudates evaluations identified a total of14 secondary metabolites and two internal signaling molecules (jasmonic and methyl-jasmonic acid) in the six tested genotypes. Some metabolites with allelopathic effect extracted from exudates were synapil alcohol, p-hydroxybenzoic acid, quercetin, and 3,5,6,7,8-penta-hydroxyflavone.
This study suggested that the long roots of canola plants could produce more allelochemicals than short roots. The biochemical analysis of canola organs and root exudates showed differences between genotypes in their production of total metabolome [47].
Allelochemicals analyzed by metabolomics in sorghum
Sorghum (Shorgum bicolor) is an important international crop with C4 metabolism that produces grain, sugar syrup, cellulose biomass, and food for human and animal consumption [48]. Sorghum is a source of phenolic acids, phenolic acid glyceride esters (caffeoyl glycerols), flavonoids (flavanones, flavonols, flavanonols and flavan-3-ol derivatives), phenylpropane glycerides, dicaffeoyl spermidine and condensed tannins (flavan-3-ols and/or flavan-3,4-diols) (figure 4). The allelopathic activity of S. bicolor has been correlated to its capacity to exudate benzoxazinoids, produced in a constitutive manner as glycoside conjugates in the plant [49]. Sorghum roots produce the quinone called sorgoleone, secreted by radicular hairs [50], and whose concentration in the ground is 10-100 μM [51]. Sorgoleone shows allelopathic activity which inhibits germination and growth of susceptible weeds at 10 μM [52]. Jandová et al. [53] characterized through UHPLC-TOF-MS (Ultra-High Performance Liquid Chromatography-Quadrupole Time-Of-Flight Mass Spectrometry) the presence of fifteen metabolites with allelopathic effects such as dipeptidic amino acids, oxylipins C18, malonil monoglycosides, as well as 2 furanocoumarins: xanthotoxin and bergap-tene. Turner et al. [48] studied sorghum metabolomics and founded high metabolic variation and determined a covariation with phenotypic traits; in particular, specific metabolite patterns influence carbon assimilation, growth during early stages, and final biomass. These authors mentioned that the potential use of early phenotype prediction through metabolome both in greenhouse and in field must be explored.

Source: own preparation.
Figure 4 Principal allelopathics metabolites ofS. bicolor. a) Sorgoleone (358.5 g/mol, C22H30O4), b) bergaptene (216.19 g/mol, C12H8O4), c) xanthotoxin (216.18 g/mol, C12H8O4), d) apigeninidin (255.24 g/mol, C15H11O4), e) luteolinidin (271.24 g/mol, C15H11O5), and f) dicaffeoyl spermidine (307.39 g/mol C16H25N3O3).
Allelochemicals analyzed through metabolomics in rye
Rye (Secale cereale) provides one of the best cover crops in the cold season. Rye is used as a result of its excellent potential for weed suppression. The main by-products of S. cereale are phenylpropanoids, in particular ferulic acids, such as 2-(E)-O-feruloilglu-conic acid found in primary leaves, or dehydrodimeric ferulic acid, as well as p-coumaric acid and synapic acid in grains. The allelopathic capacity of rye (S. cereale L.) is due to the presence of phytotoxic benzoxazinoids, which are compounds whose biosynthesis is regulated by plant development and which accumulate mostly in young tissue, as well as express dependence in crops being influenced by the environment. The discovery of natural benzoxazinoids was based on the identification of greater resistance against pathogenic fungi in rye plants. Thus, the first plant benzoxazinoids found were in rye. The compounds can be expressed from residues of rye grown in greenhouses, with levels between 12 and 20 kg/ha from reduced amounts exuded into living plants. In soil, benzoxazinoids are present in a range of 0.5 to 5 kg/ha. In addition to the research conducted on benzoxazinoids, benzoxazolin-2(3H)-one (BOA) has been explored in order to identify phytotoxic effects on weeds and crops.
Exposure to BOA affects transcriptome, proteome and metabolome patterns in seedlings. It also inhibits germination and growth and may cause the death of sensitive species [54]. The variability in sensitivity of cultivars and ecotypes results from how different species-dependent strategies for responding to BOA have evolved. Over the last few decades, the use of rye as a cover crop to control alelopathic weeds (e.g., in corn, cotton and soybean fields) has gained importance. Several flavonoids are present in young rye leaves. In addition to cyanidine glycosides, 2'-O-isovitexine glycosides, apigenine-glycosides and luteolin-O-glucuronides are found in primary leaves. While p-coumaric acid, ferulic acid and two isovitexineglycosides accumulate in the adaxial and abaxial epidermal layers, mesophilic acid contains two luteolin glucuronides. In this regard, the most important by-products are glucosylated (BX) benzoxazinoids, 2,4-dihydroxy-2H-1, 4-benzoxazin-3(4H)-one (DIBOA glucoside) in sprouts, including 2,4-dihydroxy-7-methoxy-2H-1,4-benzoxazin-3(4H)-one (DIMBOA glucoside) in roots [54] (figure 5).
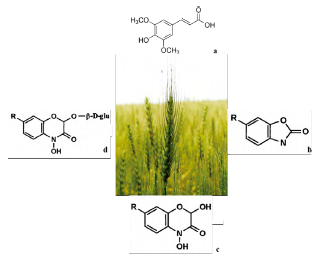
Source: own preparation.
Figure 5 Principal allelopathics metabolites of S. cereale. a) Synapic acid (224.21 g/mol, C11H12O5), b) benzoxazolin-2(3H)-one (BOA), c) (DIBOA), and d) DIMBOA glucoside.
In particular, aglycons and their derivatives are responsible for the phytotoxic effects of rye residues. However, they may act with other compounds, such as ferulic acid and its related compounds, lutein glucuronides, β-phenylacetic acid and β-hydroxybutyric acid. In particular, the tissues of young rye plants have the highest concentration of simple phenolic allelochemicals. Currently, DIBOA and BOA have been indicated as the most stable phytotoxic compounds in rye [55].
Allelochemicals analyzed by metabolomics in wheat
The accelerated development of herbicide resistance in certain weed species is a growing threat to sustainable agricultural production. A number of studies have been conducted on the management of weeds and wheat ( Triticum aestivum), since this culture is among the most representative in the world due to its high allelopathic potential. The aforementioned studies are based on a preliminary analysis of said potential and include the identification of allelochemicals and genetic markers associated with the allelopathy condition. Wheat seedlings produce and release exudates from toxic roots, which are inhibitors of several weed species [56].
In particular, Wu et al. [57, 58] mentioned that strongly allelopathic accessions of T aestivum produced significantly greater amounts of allelochemicals in young seedling shoots or roots. In addition, accessions exuded more allelochemicals into the growing medium. Two main Quantitative Trait Loci (QTL) associated with wheat allelopathy have been identified. Cloned cDNA genes may be manipulated in order to regulate the biosynthesis of allelochemicals. This mechanism might improve weed suppression through elevated levels of allelopathic potential in commercial wheat cultures. Wu et al. [57] identified significant genetic variations in the production of phenolic compounds, including benzoxazinones (a type of alkaloids), among 58 accessions of the T. aestivum variety, which were previously found with diverse allelopathic potential. Such authors mentioned, the concentration can be up to 68 times the difference in buds, occurring 41 times in roots, and 70 times in exudates of the roots. DIM-BOA (2,4-dihydroxy-7-methoxy-1,4-benzoxazin-3-one) and its compound MBOA (6-methoxybenzoxazolin-2-one) inhibited oat (Avena fatua) root growth [57, 58]. Huang et al. [59] reported that two benzoxazinones, i.e., DIMBOA and DIBOA (2,4-dihydroxy-1,4-benzoxazin-3-one) were associated with higher inhibition of the L. rigidum variety compared to natural lactama benzoxazinones, HBOA (2-hydroxy-1,4-benzoxazin-3-one) and HMBOA (2-hydroxy-7-methoxy-1,4-benzoxazin-3-one). The latter was the result of the absence of an important OH group at the fourth position of the oxazinone ring. Lavergne et al. (2018) used metabolomic methods of mass spectrometrygas chromatography (GC-MS) in order to provide a characterization of the chemical composition of cuticular waxes in wheat leaves and stems [60]. Such researchers discovered 263 putative compounds and 58 wax compounds that are part of alkanes and fatty acids. Metabolites have associations known as important biological functions. The leaves contained more primary alcohols than stems such as 6-meth-ylheptacosan-1-ol and octacosan-1-ol (figure 6).
Other crops of agroindustrial importance
Wang et al. [61] mentioned that Conyza canadensis show allelopathic effects: they discovered that leaves in C. canadensis plants secrete a higher amount of allelochemicals in high latitudes than in lower latitudes. The effects of C. canadensis on L. sativa were a decrease in plantling height, plantling biomass, germination potential, germination index, and vigor index. In sugar cane, the metabolomic approach is conducted towards mechanisms of regulation of sucrose accumulation. Thus, Bosch et al. [62] have been able to use metabolomics to examine expression profiles in metabolite levels with the use of GC-MS technology to separate and identify multiple metabolites. 30 metabolites have been identified so far in sugar cane, out of which the most important ones are amino acids and organic acids, as well as carbohydrates with an important role in signaling (for example, trehalose). The authors mentioned that the work consists in comparing the levels of metabolites in sugar cane between varieties and sugar production [62].
Biological activity
Finally, table 1 summarizes the main biological activities of the allelopathics secondary metabolites mentioned in this study.
CONCLUSIONS
The alleloPathic Phenomenon involves an ecological, chemical (isolation, identification, and characterization of allelochemicals), and physiological (interference in biochemical or Physiological Processes, both at cellular and molecular level) comPonent. Such components must be included in the study of the involvement of said phenomenon in a certain environment. The complex biochemical pathways and the distinct categories of allelopathic compounds indicate that multiple genes are probably involved in allelochemical production. Metabolomics generates genuine metabolomic profiles that allow for the study of catabolism and bioactivity of natural products. With the growing number of "omics", new approaches can be used to understand metabolic changes that occur in plants, especially in proteomics and metabolomics where literature is still scarce, and thus those analyses could be profiled in coming years with the objective of knowing the allelopathome of a larger number of plant species and their interactions. These studies suggest that a combination of secondary metabolites can be used to attack weeds. In situ and In vitro culture of secondary metabolites (allelo-chemical types) is recommended to be used as organics compounds in weed organic control in the future.
















