I. INTRODUCTION
The diagnosis of diseases is based on the recognition and analysis of clinical findings, referring to the altered signs and symptoms that the individual presents at a given time [1]. A symptom is the patient's subjective perception related to the disease, while the sign is objectively recorded by the health professional. Some manifestations are characteristic of certain diseases; in others, complementary studies are indispensable to reach a definitive diagnosis [2]. At the time of the clinical examination, the observation of the oral mucosa and skin are indicators to guide the expert to a presumptive diagnosis.
During the clinical inspection, health professionals rely on manual, electronic, and software tools to obtain an accurate diagnosis. Among the existing software tools, those based on artificial intelligence (AI) methods to simplify and improve clinical activities stand out [3]. The continuous evolution of technology has made diagnostic tools evolve and include other means to develop these activities remotely. In this context, the use of mobile devices in medical diagnostic activities is increasing, an example of which is tele-dentistry, which has boosted the use of multiple communication platforms between patients and the dentist using the phone camera to capture the lesion and process it by mobile applications [4,5]. In the current market of apps, some support both the physician and the patient, providing solutions that include ruling out symptoms through multiple questions, obtaining a possible diagnosis based on the answers received [6].
Currently, there are cases of applications that use Artificial Neural Networks (ANN) to identify lesions using a database of more than 12000 images, applications that try to diagnose through some ANN method, taking a photograph to give a medical diagnosis with a high percentage of accuracy [7]. Convolutional Neural Networks (CNN) continue to pioneer Machine Learning (ML) methods because of their fault tolerance and ease of insertion with existing technology [8]. This CNN can be configured using TensorFlow, an open-source library for training and developing ML models developed by Google©. The base unit is the tensor, which can be viewed as a multidimensional data matrix that includes image information such as width, length in pixels, and each color channel [9].
Current AI tools that support the dentist are based on the analysis of radiographic and computed tomography images [10]. However, evidence supporting the oral diagnosis of soft tissue diseases using computer vision, cell phones, and clinical images are scarce and limited. Having a mobile application that can diagnose in real-time with a simple image using smartphone hardware and software in real-time is advantageous, as it can be used to guide healthcare professionals in making a more accurate diagnosis. These tools can support those dentists who work and perform activities in rural areas where access to a specialist is difficult and complicated. In this sense, the purpose of this work was to develop a prototype mobile application for the identification of oral lesions based on convolutional neural networks.
II. METHODOLOGY
This section describes the data, techniques and methodology used for the development of the project.
A. Dataset
The dataset used in this work was constructed with images found on the internet for each disease using web scraping. 500 different images distributed in 5 classes were used as follows: 97 leukoplakia images, 101 aphthous stomatitis images, 74 nicotinic stomatitis images, 110 HSV-1 images, and 118 images from the no lesion/no recognized class were included, decreasing false positives and improving performance when classifying. To avoid overtraining of the CNN, random linear transformations such as zooms and rotations were performed on each image, increasing the dataset at least eight times from the original set. In the particular case of HVS-1 images were used to test the feature extraction function performed by the mobile CNN used.
B. Model Used
The model used for the mobile application was Mobilenet V2, as it presents the best floating-point model for mobile vision applications. This model offers a lightweight deep neural network whose architecture uses depth separable convolutions, combining this convolution with batch normalization and linear rectifier functions, additionally, softmax was used as the final classifier function. The integrated TensorFlow lite API converter generated the model compatible with mobile applications.
C. Description and Requirements of the Application
The developed mobile application has two activities and one general service. The recognition service prepares the image, loads to the CNN model, analyzes the image, performs inferences, and displays the results. The recognition model transforms the received bitmap to a tensor that is understood by TensorFlow to run the model returning the inference probability through a dynamic list with confidence percentages. Additionally, it is allowed to choose which CPU performs the inference (CPU or neural network API as hardware accelerator). Information regarding frame size, cropping, rotation, and inference time is displayed in a tab (Figure 1). The Android version targeted by the application is Android 9, the minimum being Android 4.4.2 to run and the tests were performed on a Nokia 7.1 device running Android version 9.
D. Data Preparation and Training
The data set was randomly divided into two, 80% for training and 20% for testing and validation. The preprocessing of the images, included in addition to linear transformations, adapting the image to a 640x480px RGB object. To save training and computational time, the transfer learning method of the CNN ImageNet model was applied, using Google Collaboratory and Tensorflow.
III. RESULTS
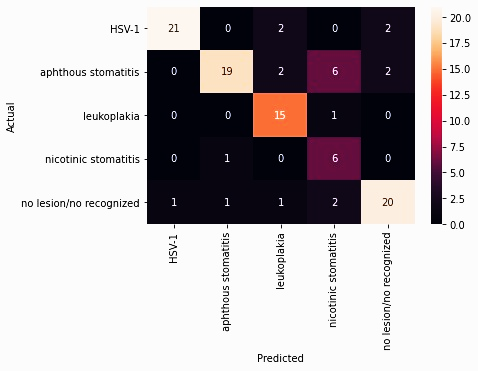
The performance and validation of the mobile application were tested with 102 images. The inference time during testing did not exceed 500ms. To establish a measurable relationship between the actual and predicted values, the confusion matrix with unbalanced data shown in Figure 2 was performed. The results of the analysis of this matrix are described in Table 1, which summarizes the findings of true positives (TP), false positives (FP), false negatives (FN), and true negatives (TN) for each class. A Kappa index of 0.739 was obtained to measure concordance.
Table 1 Results of the confusion matrix analysis.
| Class | Truth Overall | Classification Overall | TP | FP | FN | TN |
|---|---|---|---|---|---|---|
| Aphthous stomatitis | 21 | 29 | 19 | 10 | 2 | 71 |
| HSV-1 | 22 | 25 | 21 | 4 | 1 | 76 |
| Leukoplakia | 20 | 16 | 15 | 1 | 5 | 81 |
| Nicotinic stomatitis | 15 | 7 | 6 | 1 | 9 | 86 |
| No lesión | 24 | 25 | 20 | 5 | 4 | 73 |
TP: true positives, FP: false positives, FN: false negatives, TN: true negatives
The results of the confusion matrix analysis allowed the generation of metrics to compare the performance of each of the classes of the trained model (Table 2). The metrics used were accuracy, precision, sensitivity, specificity, f1-score and area under the curve (AUC) shown in Figure 3. It was observed that the HSV-1 class presented better performance in accuracy and precision, the leukoplakia class was the most sensitive in this set of tests and the nicotinic stomatitis class the most specific. To compare the overall performance between them, the AUC and f1-score metrics were used, resulting in the HSV-1 class with better performance and the nicotinic stomatitis class with unfavorable performance.
Table 2 Performance metrics for each class.
IV. DISCUSSION
The implemented model was based on CNN for the recognition of a group of oral lesions, using the Mobilenet V2 network pre-trained with ImageNet, which is characterized by being a large database with several categories including plants, flowers, animals, objects, among others, with excellent results at the time of classification [11,12]. This is the reason why this model was considered to perform the learning transfer. Lesion recognition systems using learning transfer have been described in the literature with the AlexNet [11], VGGNet [13,14], and ResNet [12,15-18] network models, whose performance is similar to the model used for this work. Regarding the training of the model, the use of TensorFlow for mobile applications of disease recognition by artificial vision has been evidenced in different research with good results [12, 16, 20, 21].
When working with images of the oral cavity, it is difficult to find many images for each case study. One of the limitations is the patients' consent for the use of their images and the uncertainty about the privacy of their data [22]. The nature of training CNN models for lesion identification requires a large image bank. In this work, the number of items per class was small, around 100 images, which involved using the method of linear transformations, zooming, and random rotations to expand the database at the time of training [9], this technique was also used by Jae-Hong et al. in their research for lesion classification in radiographic images achieving greater than 90% recognition in their classes [23].
During testing and validation, when using Tensorflow Lite, an inference time of less than 500ms was obtained, which is excellent when used as a real-time diagnostic tool. This time is considered fast for identification, as is the percentage change in certainty when the lesion is identified. The recognition of the images is not direct but made based on the probability of belonging to one of the classes programmed in the model.
The application developed presented an acceptable performance for the identification of HSV-1, aphthous stomatitis, and leukoplakia type lesions and an unfavorable performance for the recognition of nicotinic stomatitis type lesions. Unlike other applications, this one is based on clinical recognition of various oral lesions considering observation and computer vision from a smartphone. It has been evidenced that similar lesion recognition systems present a lower performance than the one obtained in this research, around 75% [14, 24-27] using image preprocessing and a higher dataset than the one used in this work. However, studies have described that good prediction performance should be greater than 90% [12, 16, 23, 28, 29]. A similar study used recognition of aphthous stomatitis-like lesions and HSV-1 with results greater than 90% using a dataset of 200 images and the random forest method as the recognition method [8].
Based on the metrics obtained the best overall performing class was HSV-1, however, in terms of accuracy and specificity attributes other classes performed better. Although the nicotinic stomatitis class obtained unfavorable and unacceptable metrics, it had the best performance in terms of specificity, this is due to the set of images that were used for training the network, despite having a sensitivity of 40% its specificity was 99%. The low sensitivity of this class was based on the similarity of the training images with other classes, which reduces the true positives recognition rate. However, when comparing these metrics with the study of A. Rana et al. it can be seen that the results of this work were superior despite having a similar data set [21].
V. CONCLUSIONS
Lesion recognition systems implemented using machine learning methods must present a good performance, to minimize the false positive and false negative rate. The developed application showed acceptable performance for three chosen lesions, however, the results reflect that this model needs improvements in the network training process. Being able to perform the recognition locally in real time using smartphones presents a great advantage for the use of this application as a diagnostic tool in remote areas without internet access.
This application would serve as an ally for dentists in early clinical diagnosis activities of potentially malignant lesions that can evolve into more complicated pathologies such as oral cancer. For this reason, further research in the area is needed to improve the features of the CNN model, such as increasing the number of images for training, separation of the clinical characteristics of the lesion to increase the rate of true positives and allowing the incorporation of other data to improve the diagnosis using additional data.