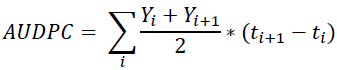Introduction
Bacterial wilt, caused by Ralstonia solanacearum, is one of the most important bacterial diseases in plants (Mansfield et al., 2012), with more than 200 host plant species belonging to over 50 different botanical families (Genin & Denny, 2011). This pathogen is present in all continents, primarily in tropical and subtropical regions, but also in Europe and North America, where cold- resistant strains are present (CABI, 2022; Janse et al., 2004; Swanson et al., 2005). Significant crop yield losses in different crops are due to R. solanacearum worldwide. For instance, up to 91% of yield losses have been reported in tomatoes, 33 to 90% in potatoes, and 70 to 100% in bananas (Elphinstone, 2005). Bacterial wilt is thus one of the most devastating plant diseases (Mansfield et al., 2012).
R. solanacearum is considered a species complex (RSSC) due to its high genetic and phenotypic diversity. For decades, the RSSC was divided into five races (R1–R5) based on the host range and six biovars (Bv1–Bv6) based on the metabolic utilization of carbon sources (Denny & Hayward, 2001). Despite being valid for many years, both systems lacked a genetic basis, became unreliable, and were replaced by the phylotype-sequevar system. DNA sequence analyses of the 16S-23S gene intergenic spacer (ITS) region, the transcriptional activator HrpB, and the endoglucanase (egl) genes allowed the classification into four major subdivisions called phylotypes (Fegan & Prior, 2005). The phylotypes correspond to the species’ geographic origin: phylotype I comprises strains from Asia, phylotype II includes two sub-clusters (IIA and IIB) from the Americas (Fegan & Prior, 2005), and phylotypes III and IV comprise strains from Africa and Indonesia, respectively (Paudel et al., 2020). Recently, Safni et al. (2014) proposed a new classification system, which distinguishes R. solanacearum into three genospecies based on phylogenetic analyses of the 16S-23S rRNA ITS gene sequences, 16S-23S rRNA intergenic space region sequences, endoglucanase sequences, and DNA-DNA hybridizations. The first genospecies includes strains from phylotype II, the second genospecies includes R. syzygii, and strains from phylotype IV. Finally, the third genospecies comprises strains belonging to phylotypes I and III (Safni et al., 2014).
Brown rot or bacterial wilt of potatoes is considered a severe threat to the potato industry. It is estimated to cause losses of 950 million dollars annually in 80 countries (Charkowski et al., 2020) and severe crop losses in tropical, subtropical, and warm temperate regions. The disease may also occur in cooler climates, such as relatively high tropic elevations or higher altitudes (Osdaghi, 2022). The bacteria invade the host through the roots, colonizing the xylem vessels in the vascular system. In the early stages of the disease, the first visible symptoms are wilting of the youngest leaves at the end of the branches during the hottest part of the day. Reduced growth, yellowing, and sudden wilt, followed by rapid death, are some later-stage symptoms (Muthoni et al., 2020).
Controlling bacterial wilt is remarkably challenging since it is a soil, water, and seed/tuber-borne pathogen and can survive in soils and water without a host (Genin & Denny, 2011). Its dissemination is a threat to crops, and it is considered a quarantine bacterium by the Animal and Plant Health Inspection Service (APHIS) of the United States Department of Agriculture (USDA), the North American Plant Protection Organization (NAPPO), the European Plant Protection Organization (EPPO) (Champoiseau et al., 2009), and the Instituto Colombiano Agropecuario (ICA, 2018). ICA was the first to report the presence of this pathogen in Colombia for potatoes in 2018 (ICA, 2018), but it was eradicated and has not been detected since then.
Controlling the presence of the pathogen in soil, water, and seeds is recommended to avoid the dissemination of this bacterium in potatoes (Karim & Hossain, 2018; Kurabachew & Ayana, 2017). Additionally, crop rotation and weed control are advised as control measures in infected soils (Ayana & Fininsa, 2017). Chemical control has not been proven to be highly efficient against bacterial wilt (Karim & Hossain, 2018). Without efficient strategies to eradicate R. solanacearum, using resistant cultivars appears to be the best strategy to control this disease in potatoes (Boschi et al., 2017).
Potato breeding for resistance against this pathogen has been done in Brazil (Brazilian Agricultural Research Corporation, Embrapa Hortaliças), Peru (Centro Internacional de la Papa, CIP), Uruguay (Instituto Nacional de Investigación Agropecuaria, INIA) and the USA (University Wisconsin). Programs have leveraged different bacterial wilt resistance/tolerance sources derived from cultivated (S. tuberosum group Phureja) and wild relative species (S. commersonii and S. chocoense) (Carputo et al., 2009; French & Lindo, 1982; Hawkes, 1994; Muthoni et al., 2020; Narancio et al., 2013; Sequeira & Rowe, 1969). However, breeding for resistance against this pathogen is complicated since the resistance/tolerance responses are highly affected by the host-pathogen-environment interactions due to their polygenic nature (Muthoni et al., 2020). Also, latent (symptomless) infections pose a significant limitation when evaluating resistance to this disease in plant germplasm. Breeders may phenotypically choose elite parentals plants that look resistant because they do not show wilting symptoms but that may bear the pathogen in latent infections (Ferreira et al., 2017; Zuluaga Cruz et al., 2013). The introgression of undesired alleles by linkage drag and the plasticity of resistance/tolerance responses (opposite responses at different temperature/high conditions) (French & Lindo, 1982; Tung et al., 1990) further complicate the breeding process. Consequently, the fixation of loci associated with R. solanacearum resistance/tolerance responses, derived mainly from wild tuber-bearing species, in cultivated potatoes is challenging (Muthoni et al., 2020).
The Colombian Central Collection (CCC) of potatoes contains around 2,069 accessions (Moreno & Valbuena, 2006), of which only 13 correspond to commercial varieties, making it a good source for plant breeding programs. Nevertheless, the collection has not been used to its full potential as a source for bacterial wilt resistance. During the 1970s, some resistant S. phureja clones found in the CCC collection were taken to the University of Wisconsin for a breeding program against bacterial wilt, intercrossing some of these clones with S. tuberosum ssp. tuberosum germplasm (Rowe & Sequeira, 1972). These clones were then sent to the CIP in Peru. For decades, the CIP breeding program relied on only two clones of S. phureja as sources of resistance; however, the resistance was very unstable, leading to the incorporation of other species (Muthoni et al., 2020). No studies regarding tolerance/resistance against R. solanacearum in the CCC collection exist. Therefore, we hypothesize that the CCC is essential for identifying tolerant/resistant genotypes against this pathogen. To this end, this study evaluated the response of 11 commercially available accessions from the CCC to R. solanacearum.
Materials and methods
Plant material
To help in the agronomical characterization of the commercially available Colombian potato cultivars, eleven (Table 1) that have not been previously characterized for R. solenacaerum were selected from the CCC of Agrosavia. Plants were in-vitro propagated using the Murashige and Skoog (MS) medium (Murashige & Skoog, 1962) supplemented with vitamins, 3% sucrose, 0.7% plant agar, at pH 5.8 and cultured in growth chambers with 16-h-light/8-h-dark photoperiod at 22 °C and average light intensity of 200 μmoles m-2.-1. Three weeks later, plants were transferred into 22 oz pots (16 cm in height and 9 cm in diameter) containing sterile soil and grown for hardening during three more weeks in a growth chamber at 22–25 °C and 50– 60% relative humidity (RH).
Bacterial strain and growth conditions
The R. solanacearum strain phylotype IIB, a highly aggressive strain isolated from plantain with typical bacterial wilt symptoms in Colombia, was kindly provided by Adriana González from Universidad Nacional de Colombia. Despite being isolated from plantain, phylotype IIB strains are known to infect highland and cold-tolerant potatoes (Otieno et al., 2021). The R. solanacearum strain was kept at -80 °C in a glycerol stock and routinely grown at 28 °C on a Triphenyl tetrazolium chloride (TTC) solid medium (Denny & Hayward, 2001) for 48 hours. To prepare the bacterial culture for inoculation, a colony from the TTC medium was taken and grown in 100 ml of nutrient broth medium at 28 °C for 24 hours in a shaker (Ohta & Hattori, 1980).
Plant inoculation and disease rating
For plant inoculation, the bacterial strain was grown in 100 ml of nutrient broth medium (Ohta & Hattori, 1980) at 28 °C with shaking at 200 rpm for 24 hours. The following day, cells were pelleted by centrifugation, suspended in water, and adjusted using a spectrophotometer (Thermo Scientific Genesys 30) to 1x10. CFU/ml (OD600 = 1 for R. solanacaerum). Six-week-old plants were drench-inoculated with 40 ml of 1x10. CFU/ml bacterial suspensions. Control plants were mock-inoculated with 40 ml of water.
Eight plants per genotype were inoculated for disease rating in a complete randomized design in three independent experiments. After inoculation, plants were maintained in a growth chamber at 22–25 °C (50–60% RH) with a 12-h photoperiod. Disease symptoms were first observed two weeks after inoculation. Development was recorded every two days or up to 25 days after inoculation using an ordinal disease index scale: 0 (no wilting symptoms), 1 (0–25% of the leaves showing wilting symptoms), 2 (25– 50% of the leaves showing wilting symptoms), 3 (50–75% of the leaves showing wilting symptoms), 4 (75–100% of the leaves showing wilting symptoms), and 5 (all leaves wilted and plant dead) (Abdrabouh et al., 2019). The area under the disease progress curve (AUDPC) was calculated 15 to 25 days after inoculation (Pedroza & Samaniego, 2009).
Y: Disease index value
t: Test day
The AUDPC data from the three replicated experiments were analyzed by a Generalized Linear Model (GLM). The differences between the means were calculated by LSD Fisher’s test with Bonferroni correction. Additionally, using the disease index (DI), the phenotypic response of each genotype was characterized as resistant or susceptible based on Gorshkov and Tsers (2022); DI = 0 was established as resistant and DI > 0 as susceptible.
Results and Discussion
Disease evaluation showed that six accessions, including five S. tuberosum group Andigena cultivars (Diacol Capiro, Ica Unica, Mary, Parda Fina, and Parda Pastusa) and one from S. tuberosum group Phureja (Criolla Colombia) were highly susceptible to this pathogen with values ranging from 3.62 to 4 (Table 1). In contrast, five accessions, including one S. tuberosum group Phureja (Sol Andina) and four S. tuberosum group Andigena (Pastusa Superior, Perla Negra, Roja Nariño and Superior), were highly resistant/tolerant (Table 1).
Table 1. Disease index (DI) and classification of each potato genotype evaluated as resistant (DI= 0) or susceptible (DI > 0)
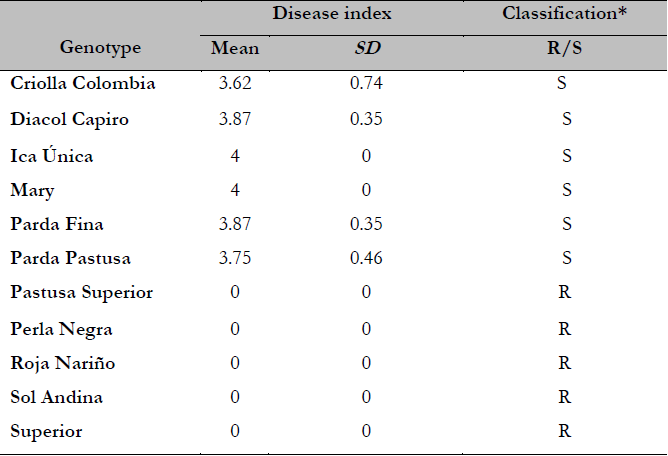
* ClassificationResistant (R); Susceptible (S).
Source: Prepared by the authors
Bacterial wilt symptoms of three susceptible and three resistant accessions are depicted in Figure 1. Phenotypic evaluation of the potato accessions shows that susceptible accessions are completely wilted 25 days after inoculation (Figure 1A), while resistant varieties lack symptoms (Figure 1B).

Source: Prepared by the authors.
Figure 1. Phenotypic evaluation of potato germplasm for resistance against R. solanacearum. Symptoms are developed in three susceptible accessions: Mary, Ica Unica, and Parda Pastusa (A), compared to symptomless resistant accessions: Sol Andina, Superior, and Roja Nariño (B) at the end of the experiment, 25 days after inoculation. The vertical white line corresponds to 16 cm.
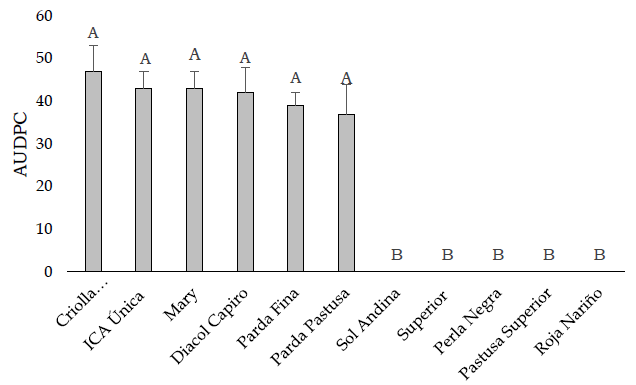
The aggressiveness of R. solanacaerum, calculated using the AUDPC, was evaluated in all accessions from the susceptible cultivars: the accessions of S. tuberosum group Phureja (Criolla Colombia) and S. tuberosum group Andigena (Diacol Capiro, Ica Unica, Mary, Parda Fina ,and Parda Pastusa) showed a similar degree of susceptibility against the R. solanacearum isolate (Figure 2).
These results are relevant for Colombia since it implies that there are strains in the field capable of causing disease on the potato cultivars that the farmers currently use. Also highlights the importance of having germplasm banks to search for resistance/tolerance traits. In the case of Colombia, five highly resistant accessions against this pathogen are commercially available. Still, there was a lack of knowledge of these resources since these accessions were not characterized before for this pathogen.
Source: Prepared by the authors.
Figure 2. Aggressiveness of R. solanacearum on potato genotypes. Mean area under the disease progress curve (AUDPC) values for three experiments with eight replicates and standard deviation of the genotype are depicted. The resistant accessions exhibited symptoms at the end of the experiment, 25 days after inoculation. AUDPC data were subjected to GLM analyses in R 3.6 using the lme4 package (Bates et al., 2019). The capital letters significantly indicate differences (p ≤ 0.05) among genotypes according to LSD Fisher’s test with Bonferroni correction.
Breeding for resistance against bacterial wilt is probably the best option to control this disease. The use of antibiotics in agriculture is banned almost worldwide, chemicals are not efficient, biological control agents are not commercially available, and cultural practices are difficult to apply (Boschi et al., 2017; Champoiseau et al., 2009). Thus, finding these five tolerant accessions with agronomically desirable traits is a significant step towards having a reservoir for bacterial wilt resistance cultivars ready to be deployed to the field. Also, they could be used as resistant parentals in breeding programs. It is important to note that resistance genes have been previously found in wild potato species, such as Solanum commersonii, S. stenotomum, and S. goniocalyx, among others; however, resistance was affected by high temperatures, and the plants had latent infections (Ferreira et al., 2017; González et al., 2013; Narancio et al., 2013; Zuluaga Cruz et al., 2013). Resistant clones of S. commersonii have reduced wilting symptoms rather than complete immunity against R. solanacearum (Otieno et al., 2021; Zuluaga Cruz et al., 2013). Because the five tolerant accessions found in this work did not show wilting symptoms 25 days after inoculation, it will be a natural next step to determine the nature of the resistance, whether it is due to a quantitative trait locus (QTL) or R-genes involved. It will be critical to determine whether these resistant cultivars can harbor bacteria in latent infections, select parental cultivars with the potential for plant breeding programs and understand the resistance’s nature. However, these results show a potential added value in some of these commercial varieties for potato growers and plant breeders. These results highly impact Colombian agriculture since new sources of resistance against this devastating pathogen in rarely used commercial varieties ready to be deployed on the field were found.
Conclusions
The Colombian Potato Germplasm Bank constitutes an essential source of genetic diversity, and screening these genotypes against potato bacterial wilt is necessary to identify resistant varieties. To the best of our knowledge, this is the first study characterizing 11 potato accessions against R. solanacearum commercially available in Colombia, demonstrating the importance of germplasm banks as a source for plant breeding programs to introgress important agronomical traits.