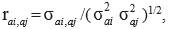Introduction
Estimates of genetic and phenotypic parameters for growth traits of Mexican beef cattle (e.g., Limousin, Charolais, Charbray, Simmental, Simbrah, Indubrazil) are available in the scientific literature (Ríos-Utrera et al., 2011; Ríos-Utrera et al., 2012; Vega-Murillo et al., 2012; Ríos-Utrera et al., 2013). Despite several Mexican breeders associations conduct national cattle evaluations of scrotal circumference as an indicator of male fertility (sperm volume and quality), and frame score as a measure of lean-to-fat ratio potential, only one study (Torres-Vázquez et al., 2012) has estimated genetic and phenotypic parameters for scrotal circumference (SC), frame score (FS), and yearling weight (YW), and their relationships in Mexican beef cattle.
Multivariate analysis reduce the prediction error variance and in some instances reduce or eliminate bias from selection (Schaeffer, 1984). Additionally, knowledge of genetic relationships among economically important traits helps to decide which traits to include as selection criteria when developing breeding selection objectives and to predict the net impact of selection. Scrotal circumference of yearling bulls is moderately heritable (Koots et al., 1994). Its genetic correlation with age at first calving (Barrozo et al., 2012) and pregnancy rate in female relatives is favorable (Eler et al., 2006). In addition, selection for greater SC may simultaneously increase postweaning growth rate, YW, weight at 18 months of age and FS. Some researchers have reported positive genetic correlation estimates for SC with those traits (Johnson et al., 1993; Crews and Porteous, 2003). The objective of this study was to estimate heritabilities and genetic, environmental, and phenotypic correlations, and direct (DRS), and correlated (CRS) responses to selection of scrotal circumference (SC), frame score (FS), and yearling weight (YW) of Mexican Charolais (CH), and Charbray (CB) young bulls.
Materials and Methods
Population
The study involved Charolais and Charbray cattle breeds. Charolais included upgraded (31/32 Charolais-1/32 Brahman) and fullblood cattle. Charbray consisted of 5/8 Charolais and 3/8 Brahman. The Charolais-Charbray Herd Book of Mexico (CCHBM) provided productive and pedigree data used for the study. Records were collected on a total of 10,078 Charolais and 500 Charbray young bulls, born from 2004 to 2015. The bulls were the progeny of 1,741 sires and 8,755 dams. The pedigree file consisted of 20,906 animals. Connectivity among different Charolais and Charbray herds has been established to some extent through the use of common semen of Charolais and Charbray sires from U.S.A., France and Mexico, and through auction of bulls and heifers among Mexican breeders.
Measurements
The traits analyzed were SC, FS and YW. Qualified CCHBM technicians measured SC and height of the bulls. Scrotal circumference was measured in centimeters at the widest part of the testis using a flexible tape. Animal height and age records were used to calculate FS. The original database consisted of 28,299 records. The age range of the animals was 320 to 410 d for the three traits; therefore, records outside these ranges were eliminated from the analysis, but the animals remained in the pedigree file. Productive data was edited to remove unreliable SC, FS and YW measurements (± 3 standard deviations from the mean) as well as unreliable dates. In addition, contemporary groups with only one bull were eliminated. The pedigree file was checked to make sure all parents were born before their progeny. Consequently, 17,721 records (62.6%) were excluded from the original database. Actual SC, height and YW records were adjusted to a standard animal age of 365 d. Because the CCHBM had not developed their own age adjustment factors for SC, the 0.0505 adjustment factor for Charolais cattle recommended by the Beef Improvement Federation (BIF, 2002) was used to calculate adjusted 365-d yearling SC for each bull of the pure and the composite breeds evaluated. The SC was adjusted with the following formula: SC=Yearling SC + [(365 - Age) x 0.0505]. Adjustments for YW and FS followed the Guidelines for Uniform Beef Improvement Programs, published by the Beef Improvement Federation (BIF, 2002). We analyzed FS instead of hip height because FS is easier to interpret and more applicable than hip height. Table 1 presents descriptive statistics for SC, FS, and YW. For Charolais and Charbray bulls, raw means of SC, FS and YW were 29.9 ± 2.8 and 28.7 ± 3.4 cm, 5.0 ±1.2 and 5.5 ± 1.2 units, and 347.6 ± 48.7 and 332.4 ±42.6 kg, respectively.
Model
A three-trait animal model was fitted to estimate variance and covariance components and genetic parameters. The animal model for each trait included breed of bull, contemporary group, age of dam in days as a linear covariate, direct additive genetic effect and residual. Contemporary group was defined as a group of young bulls born in the same herd, year, and season. The season effect included four classes: January- March, April-June, July-September and October- December. The number of contemporary groups for SC and FS was 1,812; for YW, however, the number of contemporary groups was 1,919. The number of contemporary groups for YW was different to that for SC and FS, because YW measurements were not taken the same day as SC and FS measurements. For all traits, the smaller contemporary group had two bulls, while the larger contemporary group had 58 bulls; however, mean numbers of bulls per contemporary group were 5.5 for YW, and 5.8 for SC and FS.
Table 1 Summary statistics for scrotal circumference (SC), frame score (FS), and yearling weight (YW) of young bulls.
| Trait/breed | N | Mean | Min | Max | SD | CV (%) |
|---|---|---|---|---|---|---|
| Scrotal circumference (cm) | ||||||
| Charolais | 10,078 | 29.9 | 21.7 | 40.1 | 2.8 | 9.4 |
| Charbray | 500 | 28.7 | 17.9 | 36.3 | 3.4 | 11.8 |
| Both breeds | 10,578 | 29.8 | 17.9 | 40.1 | 2.8 | 9.4 |
| Frame score (units) | ||||||
| Charolais | 10,078 | 5.0 | 1.0 | 9.0 | 1.2 | 24.0 |
| Charbray | 500 | 5.5 | 0.5 | 9.1 | 1.2 | 21.8 |
| Both breeds | 10,578 | 5.1 | 0.5 | 9.1 | 1.2 | 23.5 |
| Yearling weight (kg) | ||||||
| Charolais | 10,078 | 347.6 | 135.0 | 577.8 | 48.7 | 14.0 |
| Charbray | 500 | 332.4 | 223.8 | 464.3 | 42.6 | 12.8 |
| Both breeds | 10,578 | 346.9 | 135.0 | 577.8 | 48.6 | 14.0 |
N= number of records; Min= minimum value; Max= maximum value; SD= standard deviation; CV= coefficient of variation.
The three-trait animal model in matrix form was as follows:
Where:
y1, y2 and y3 are vectors of single trait phenotypic observations of SC, FS and YW.
β1, β2 and β3 are vectors of fixed effects (breed of bull, contemporary group, age of dam).
a1, a2 and a3 are unknown vectors of random direct additive genetic effects.
e1, e2 and e3 are unknown vectors of random temporary environmental effects.
X1, X2 and X3 are known incidence matrices relating phenotypes measured with fixed effects in β1, β2 and β3, respectively.
Z1, Z2 and Z3 are known incidence matrices relating phenotypes measured with additive genetic effects in a1, a2 and a3, respectively.
The first and second moments were assumed to be
Where:
G is the matrix of genetic variances and covariances.
? is the right direct product operator or Kronecker product.
A is the matrix of additive genetic relationships among animals, including foundation animals without records.
R is the matrix of residual variances and covariances.
I is an identity matrix of order equal to the number of animals with records. In addition,  ,
,  and
and  are the additive genetic variances for traits 1, 2 and 3, respectively.
are the additive genetic variances for traits 1, 2 and 3, respectively.  ,
,  and
and 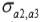 are the genetic covariances between traits 1 and 2, 1 and 3, and 2 and 3, respectively.
are the genetic covariances between traits 1 and 2, 1 and 3, and 2 and 3, respectively.
 ,
,  and
and  are the residual variances for traits 1, 2 and 3, respectively.
are the residual variances for traits 1, 2 and 3, respectively.
 ,
,  and
and  are the residual covariances between traits 1 and 2, 1 and 3, and 2 and 3, respectively. Traits evaluated were assumed to follow a multivariate normal distribution.
are the residual covariances between traits 1 and 2, 1 and 3, and 2 and 3, respectively. Traits evaluated were assumed to follow a multivariate normal distribution.
Software, initial values and convergence
Variance and covariance components, as well as genetic correlations, for genetic and temporary environmental effects were estimated with the MTDFREML package (Boldman et al., 1995). Estimates of variance and covariance components from the scientific literature were used as initial values in preliminary analyses (one for each response variable) fitting single-trait animal models. Estimates of variance and covariance components from single- trait analyses were used as priors for the three-trait analysis. The convergence criteria (minimum variance of the function values, -2 log likelihood) were set to 1 x 10-10 in each type of analysis. Iterations were checked to converge at a global maximum rather than to a local one. Results from the first run were used as starting values for up to six cold restarts to ensure convergence to the same solutions.
Description of genetic, residual and phenotypic parameters, and response to selection
Heritability for direct additive genetic effects 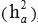 , and genetic (rai,aj), residual (rei,ej) and phenotypic (rpi,pj) correlations between traits i and j were estimated as follows:
, and genetic (rai,aj), residual (rei,ej) and phenotypic (rpi,pj) correlations between traits i and j were estimated as follows:
Where:
 is the phenotypic variance, which was estimated as
is the phenotypic variance, which was estimated as 
 is the phenotypic covariance between traits i and j.
is the phenotypic covariance between traits i and j.
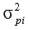 is the phenotypic variance fortrait i.
is the phenotypic variance fortrait i.
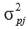 is the phenotypic variance fortrait j.
is the phenotypic variance fortrait j.
Remaining estimates of variance and covariance were previously defined.
Expected direct response to selection (DRS) was obtained for SC, FS and YW. Expected correlated response to selection (CRS) was obtained for SC and FS from indirect selection on YW, because Mexican Charolais and Charbray breeders use to select bulls based on YW. Responses to selection (one generation away) were calculated from selection of sires based on half-sib progeny records, using the formulas of Van Vleck et al. (1987):
Where:
P is the number of half-sib progeny (5, 10, 20, 50, 100, 200, and 500).
λ is equal to (4- h x 2 )/ h x 2 .
i is the intensity of selection, which was assumed to be 1.
hx is the square root of the heritability of trait x ( hx 2 ).
σx is the phenotypic standard deviation of trait x.
r g is the genetic correlation between traits x and y.
hy is the square root of the heritability of trait y.
σ py is the phenotypic standard deviation of trait y.
Results
Estimates of genetic and residual variance and covariance are presented in Table 2.
Table 2 Genetic (lower triangular), and residual (co)variance (upper triangular) for scrotal circumference (SC), frame score (FS), and yearling weight (YW).
| SC | FS | YW | |
|---|---|---|---|
| SC | 2.55 | 0.40 | 16.80 |
| 0.67 | |||
| FS | 0.04 | 0.38 | 6.07 |
| 0.13 | |||
| YW | 4.60 | 2.31 | 582.28 |
| 235.90 |
Estimates of phenotypic variance, phenotypic covariance, and phenotypic correlations for SC, FS, and YW are shown in Table 3.
Table 3 Phenotypic variance (on diagonal), phenotypic covariance (below diagonal), and phenotypic correlation (above diagonal) for scrotal circumference (SC), frame score (FS), and yearling weight (YW).
| SC | FS | YW | |
| SC | 3.21 | 0.34 | 0.42 |
| FS | 0.44 | 0.51 | 0.41 |
| YW | 21.40 | 8.37 | 818.18 |
Estimates of heritability, genetic correlations and residual correlations for the same traits are presented in Table 4.
Table 4 Heritability (on diagonal), genetic correlation (below diagonal), and residual correlation (above diagonal) for scrotal circumference (SC), frame score (FS), and yearling weight (YW).
| SC | FS | YW | |
| SC | 0.21 ± 0.04 | 0.40 ± 0.04 | 0.44 ± 0.05 |
| FS | 0.15 ± 0.14 | 0.25 ± 0.04 | 0.41 ± 0.05 |
| YW | 0.37 ± 0.16 | 0.42 ± 0.16 | 0.29 ± 0.04 |
The estimates of heritability for SC, FS and YW were 0.21 ± 0.04, 0.25 ± 0.04 and 0.29 ± 0.04, respectively. The estimates of the genetic correlations for SC with YW (0.37 ± 0.16) and for FS with YW (0.42 ± 0.16) were similar to corresponding estimates of phenotypic correlations (0.42 and 0.41); however, the estimate of the genetic correlation for SC with FS (0.15 ± 0.14) was less than half the corresponding phenotypic correlation estimate (0.34). In general, estimates of residual correlations for SC with FS (0.40± 0.04), SC with YW (0.44 ± 0.05), and FS with YW (0.41 ± 0.04) were similar to the analogous estimates of phenotypic correlations.
For SC and FS, direct responses to selection were more than two-fold greater than the corresponding correlated responses to selection, independently of the number of half-sib progeny. Direct and correlated responses to selection based on 500 half-sib progeny were about two-fold greater than those based on 5 half-sib progeny. Correlated responses to selection based on 100, 200 and 500 half-sib progeny were similar for SC, as well as for FS (Table 5).
Table 5 Expected direct (DRS), and correlated responses to selection (CRS) from selection of sires based on different number of half-sib progeny.
| DRS | CRS | ||||
|---|---|---|---|---|---|
| Number of progeny | YW | SC | FS | SC | FS |
| 5 | 8.17 | 0.38 | 0.18 | 0.16 | 0.08 |
| 10 | 10.20 | 0.49 | 0.23 | 0.20 | 0.10 |
| 20 | 12.03 | 0.60 | 0.27 | 0.24 | 0.12 |
| 50 | 13.75 | 0.70 | 0.31 | 0.27 | 0.13 |
| 100 | 14.50 | 0.76 | 0.33 | 0.29 | 0.14 |
| 200 | 14.93 | 0.79 | 0.34 | 0.29 | 0.15 |
| 500 | 15.21 | 0.81 | 0.35 | 0.30 | 0.15 |
Discussion
Scrotal circumference heritability
Scrotal circumference was medium heritable in the Mexican Charolais and Charbray cattle population, suggesting that this fertility trait may be changed by direct selection. However, response to selection would be slow. The heritability estimate for SC reported in the present study is similar to estimates reported by other authors. In particular, the estimate for SC obtained by Everling et al. (2001) for Angus-Nelore crossbred cattle with different breed composition is similar to our estimate. In a more recent study carried out in Brazil, Boligon et al. (2006) found 0.22 heritability estimate. Gressler et al. (2000) reported that SC was 24% heritable in Nelore cattle. In South Africa, Marle-Koster et al. (2000) obtained 0.25 heritability estimate for Hereford cattle. In contrast, Gargantini et al. (2005), in several beef breeds from the Roman L. Hruska U.S. Meat Animal Research Center, found 0.05 heritability estimate for SC. On the other hand, other studies have reported moderate to high heritability for SC. Arthur et al. (2001), Dias et al. (2003), Van Melis et al. (2010), Barrozo et al. (2012) and Corbet et al. (2013) reported 0.43 ± 0.06, 0.35 ± 0.03, 0.48 ± 0.01, 0.53 ± 0.04, and 0.65 ± 0.08 heritability estimates, respectively. These values, along with those previously mentioned, including our estimate, reveal great variation in heritability estimates for SC. This variation could be due to different environments, breeds, selection intensity, effects included in the animal model, and type of models (sire model vs animal model; univariate vs bivariate model) among studies (Koots et al., 1994).
Frame score heritability
Similar to SC, FS was medium heritable, suggesting that genetic progress from direct selection on FS adjusted to 365 d of age might be slow as well. Our heritability estimate for FS is similar to the estimate reported by Horimoto et al. (2006) for Nelore cattle raised in Brazil (0.23). However, Johnson et al. (1993) and Torres-Vázquez et al. (2012) found that FS was moderately heritable (0.42) in American Brangus and Mexican Simmental and Simbrah cattle. Except for these three estimates, no other heritability estimates for this trait were found in the literature. In general, heritability estimates for hip height (0.60, 0.46, 0.48) reported in the literature (Afolayan et al., 2007; Yokoo et al., 2010; Regatieri et al., 2012) are higher than our FS estimate. Although FS derives from hip height, this comparison may not be appropriate due to significant loss of variability when a continuous variable (e.g., hip height) is transformed into a categorical variable (e.g., FS), resulting in smaller heritability estimates (Mercadante et al., 2004).
Yearling weight heritability
Yearling weight was a little more heritable than SC and FS in the present study. The current estimate for YW also indicates that this growth trait would respond satisfactorily to direct selection. Our heritability estimate for YW (0.29) is comparable with 0.33 (Crossbred cattle), 0.31 (Canchim cattle), and 0.26 (Nellore cattle) reported by Afolayan et al. (2007), Mucari et al. (2007) and Boligon et al. (2010), respectively. Our figure was smaller than estimates (0.50, 0.38) reported by Bergen et al. (2005) and Schiermiester et al. (2015), respectively, for Canadian and American crossbred cattle. Moreover, it was greater than the Pico et al. (2004) and Assan and Nyoni (2009) estimates (0.14, 0.18), respectively. The literature review by Koots et al. (1994) reported 0.33 and 0.35 weighted and unweighted heritability means for YW, respectively, similar to our estimate.
Genetic correlation between SC and FS
The low genetic correlation estimate between SC and FS found in our study indicates that genes involved in the expression of both traits are different. Thus, direct selection for SC would not affect FS. In agreement with this result, the genetic correlation estimate between both traits reported by Johnson et al. (1993) was also low (0.20). In contrast, the estimate by Torres-Vázquez et al. (2012) for the Mexican Simmental-Simbrah population was high (0.59). No other estimates of this correlation were found in the scientific literature.
Genetic correlation between SC and YW
Our genetic correlation estimate between SC and YW was moderate and positive, suggesting that there would be an increment in YW if emphasis is placed on selection to enhance SC. Other literature reports also indicate that this genetic correlation is moderate and positive. Crews and Porteous (2003), in Canadian Hereford bulls, and Torres-Vázquez et al. (2012) in Simmental and Simbrah bulls, found the same figures as we did (0.38 and 0.36 genetic correlation estimates for SC and YW, respectively). The estimate reported by Garnero et al. (2001), working with Nelore cattle in Brazil, was 0.40. Johnson et al. (1993), Marle- Koster et al. (2000) and Frizzas et al. (2009) also estimated positive genetic correlations for SC and YW in Brangus (0.23), Hereford (0.17) and Nelore (0.21) cattle, but their estimates were lower. In the analysis of published genetic correlation estimates carried out by Koots et al. (1994), the weighted mean estimate was 0.39 for SC and YW. This result closely agrees with our estimate. The study by Mwansa et al. (2000) demonstrated that a two-trait animal model including SC and concomitant body weight would result in more accurate predictions (smaller standard errors of prediction) of expected breeding values for SC than a single-trait animal model.
Genetic correlation between FS and YW
Similar to the genetic correlation estimate found between SC and YW, the moderate and positive estimate for the genetic correlation between FS and YW revealed the presence of pleiotropic gene effects. This means that genes that control FS also control YW; therefore, yearling bulls that excel in FS should also excel in YW. The genetic correlation estimate for FS and YW in our study is very similar to 0.43 and 0.47 estimates reported by Horimoto et al. (2006) and Torres-Vázquez et al. (2012), respectively, but not as strong as the 0.67 reported by Johnson et al. (1993). We did not find other published estimates for this correlation.
Expected responses to selection
Expected direct responses to selection were about two-fold greater than expected correlated responses to selection for SC and FS from indirect selection on YW. Van Vleck et al. (1987) reported that situations in which indirect selection is more effective than direct selection are rare.
In conclusion, estimates of heritability for SC, FS and YW were low, but YW tended to be more heritable than SC. Genetic correlation estimates for YW with SC and FS were moderate, but genetic correlation estimate for SC with FS was low. Direct selection for YW is expected to be effective. However, indirect selection for SC and FS based on YW would not be expected to be as effective as direct selection for improving SC and FS.